Small Molecule Ligands
What is a ligand?
Many biological molecules interact with small molecules, such as cofactors, metabolites, or drugs, collectively defined as “ligands” (Figure 1). wwPDB has also defined a larger collection of small molecules and molecular subunits, termed “chemical components,” that includes ligands as well as the standard building blocks of biological molecules. wwPDB has adopted a consistent scheme for representing and archiving this complex collection of molecules for use in the PDB archive.
In the PDB archive, every chemical component is assigned an identifying code. This includes both standard amino acids (i.e., the 20 amino acids commonly found in proteins) and non-standard amino acids (e.g., hydroxyproline) or nucleotides (guanine) or ligands (glucose). For example, the ID code for alpha D-glucose is “GLC,” and the code for the heme cofactor found in proteins like myoglobin is “HEM”. These codes identify the chemical components in PDB entry files, and can be used to search and explore information on the RCSB PDB website.
Detailed information about each chemical component is stored in the wwPDB Chemical Component Dictionary (CCD). Each definition contains descriptions of chemical properties such as stereochemical assignments, chemical descriptors (SMILES and InChI), systematic chemical names, chemical connectivities and idealized coordinates (generated using Molecular Networks' Corina, and if there are issues, OpenEye's OMEGA). These chemical descriptors uniquely define each type of ligand, based on its chemical composition and pattern of covalent bonding between atoms. This is useful for identifying all of the instances of a particular ligand in the PDB. For example, many entries include ATP in different conformations with different coordinates, but all of these share the same chemical composition and covalent structure.
Each chemical compound found in the Chemical Component Dictionary has a corresponding Summary page that highlights this information integrated with information from DrugBank about medical or health applications of the molecule. For an example, explore Tylenol, which has the CCD ID TYL.
Representation of Ligands in Entry Files
PDB entry files include a variety of information that define the characteristics of the ligand, its interactions with the macromolecule, and coordinates of the atoms. The format of this information is a bit different in the mmCIF and PDB formats, but the information included is similar. More information on specific formats is published at the wwPDB.
Chemical information represented in PDB entries includes the number of atoms in the ligand, formal name of the ligand and synonyms (if any), residue name used for the ligand, and chemical formula.
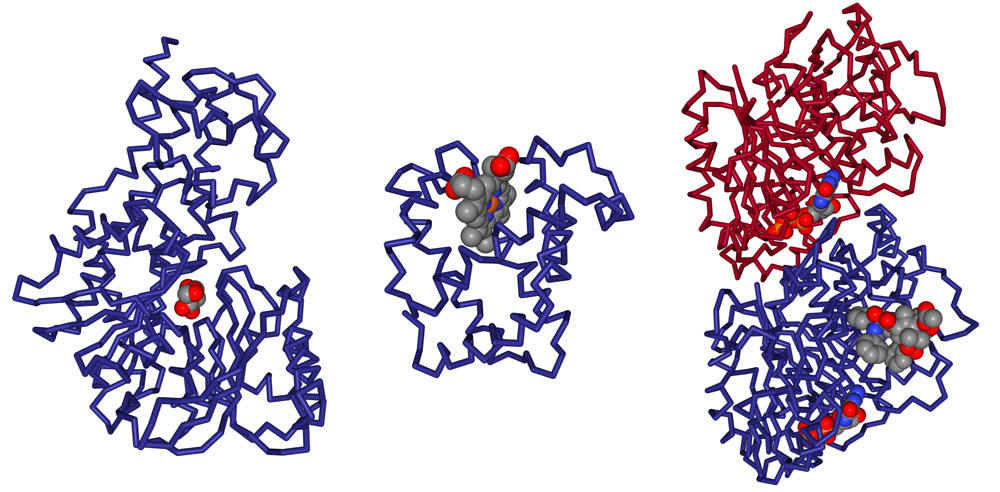
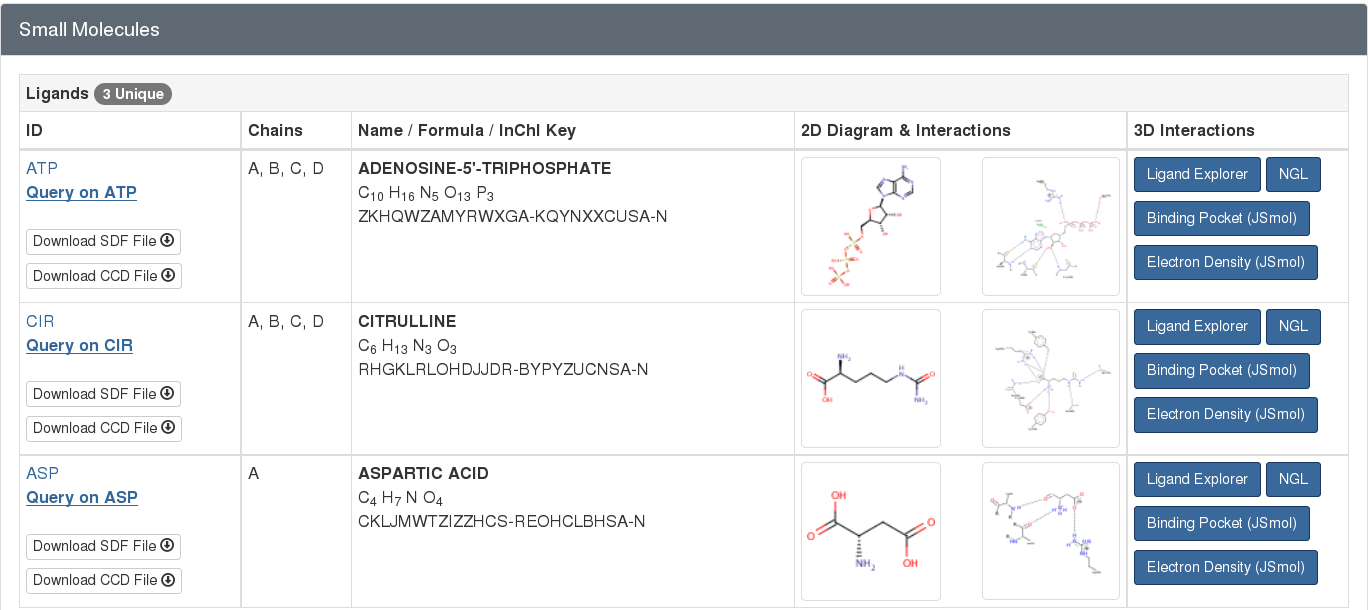
For example, the PDB entry 1j1z has 3 small molecules complexed with the protein (as seen in Figure 2). This information is highlighted from the “Small Molecule” section of the entry’s Structure Summary page at rcsb.org.
Free Ligands vs. Ligands in Polymers
Most ligands are considered “free ligands” that interact non-covalently with macromolecules. Less frequently, “ligands in polymers” found in the PDB archive form covalent bonds with macromolecules, or are included as part of polymers. PDB data are annotated to represent these different cases clearly.
Standard amino acids or nucleotides, which are normally found in protein or nucleic acid polymers, may also be found in the PDB archive as free ligands. For example, the standard amino acid aspartate (ASP) is a free ligand in the structure of the argininosuccinate synthetase (PDB entry 1j1z; Figure 2). Since this aspartate is not acting in its typical role as a component of a protein chain, it is represented as a ligand within the entry.
Conversely, sometimes ligands that are usually found in an unbound state may be included as part of a polymer chain. The most common examples are “modified” forms of the “standard” amino acids or nucleotides. For example, the modified amino acid hydroxyproline (HYP) is found as a major component of collagen as seen in PDB entry 1cag. Similarly, many proteins are engineered with selenomethione (MSE) to assist with structure determination by X-ray crystallography. In these cases, the ligand is represented as part of the protein chain and as a non-standard amino acid.
When a free ligand has covalent or metal coordination interaction with other residues this interaction is reflected in an entry’s data file. Connectivity between residues that is not implied by the primary structure is recorded. For example, in PDB entry 1pwc, penicillin G (open form) forms a bond with the catalytic serine in the bacterial enzyme DD-peptidase. This covalent interaction can be visually seen by selecting the NGL viewer button for the ligand PNM in the entry (Figure 3). PNM (in ball-and stick representation) is covalently bound to Ser62 (shown in stick representation).
Biologically Interesting Molecules
A special resource, the Biologically Interesting molecule Reference Dictionary (BIRD), has been developed by the wwPDB to classify complex ligands that are composed of several subcomponents connected in a specific way. These include peptide-like inhibitors and complex antibiotics, ribosomally synthesized gene products, such as thiostrepton (found in PDB entry 1e9w), and products of nonribosomal enzymatic synthesis, such as vancomycin (found in PDB entry 1sho). BIRD is an external reference dictionary (similar to the Chemical Component Dictionary) that provides information about the chemistry, biology, and structure of complex ligands and peptide-like-molecules. BIRD entries include molecular weight and formula, polymer sequence and connectivity, descriptions of structural features and functional classification, natural source (if any), and external references to corresponding UniProt or Norine entries. Similarly, the antibiotic Viomycin (PRD_000226; Figure 4) can be best represented as a polymer with the sequence (KBE DPP SER SER UAL 5OH).

The same BIRD molecule will have uniform representation across the archive. An important feature of BIRD is that both sequence and chemical information are provided, regardless of whether the molecule is represented as a polymer or as a ligand in the PDB archive. BIRD is regularly reviewed for consistency and accuracy, and is used to uniformly annotate PDB entries containing these molecules. The dictionary is updated each week with new definitions as the corresponding PDB entries are released in the PDB archive. The corresponding BIRD ID code appears in the PDBx/mmCIF-formatted entry file, and is used for searching and reporting at rcsb.org.
Electron Density Visualization
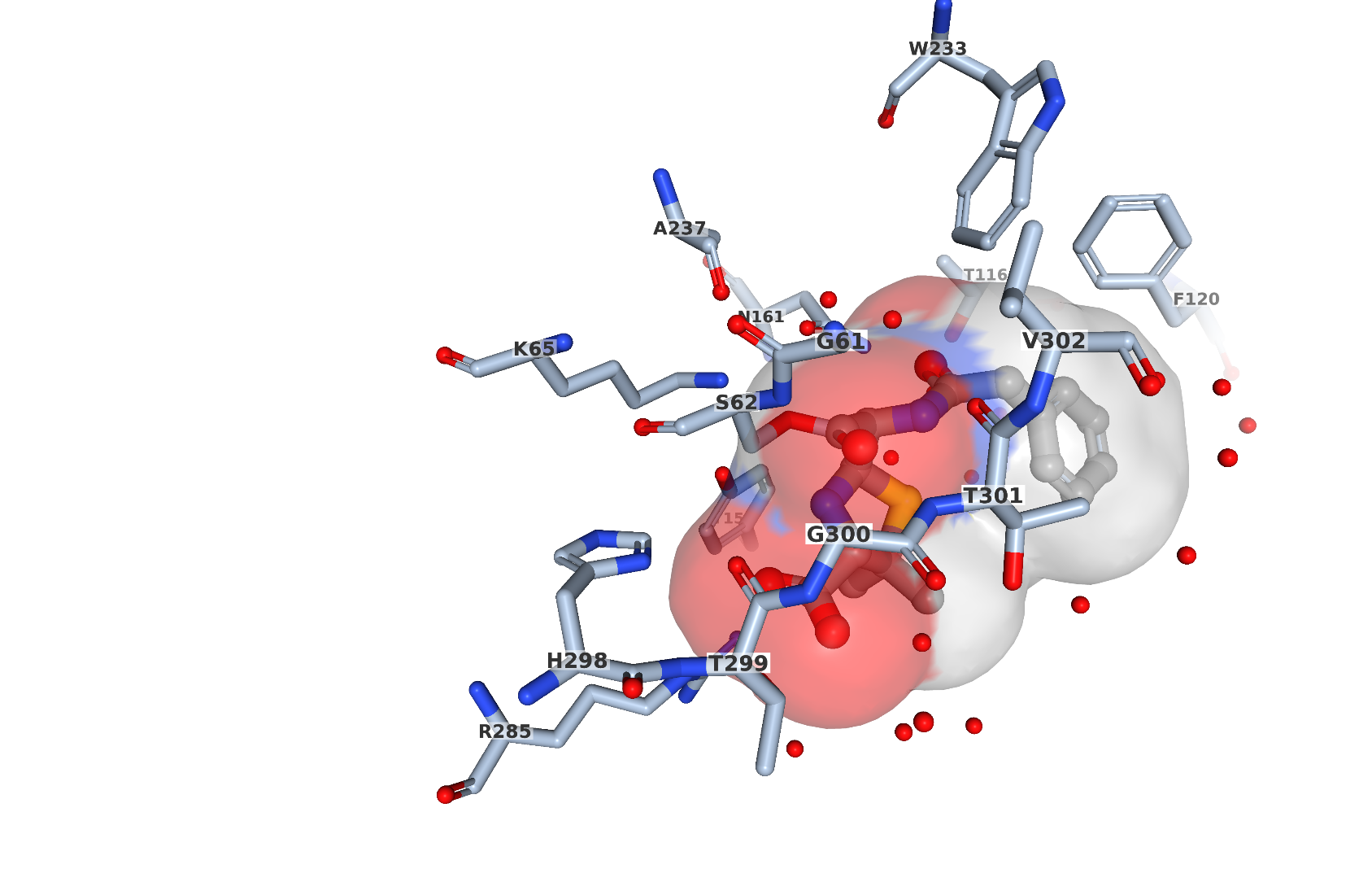
For X-ray crystal structures, the electron density defining the ligand position may also be visualized in JSmol, through the button “Electron Density” (Figure 5). This displays a crystallographic sigma-weighted 2m|Fo|-d|Fc| electron density “mini-maps” for bound ligands using the JSmol 3D View. This feature is available for ligands with more than one atom (ions excluded) in PDB entries with structure factor data. Different display thresholds (sigma values) can be selected (default threshold is 1 sigma). The mini-map, obtained from the corresponding experimental data, shows the electron density map (blue mesh, contoured at 1σ level), and represents the modeled position of the ligand (represented as ball-and stick-model) in the structure. This is useful for evaluating the experimental support for a particular ligand structure. For instance, in entry 1pwc, there is continuous electron density (Figure 5) between C7 atom of the ligand and oxygen of serine 62 providing the evidence of covalent linkage between the protein and the ligand (PNM which is open form of penicillin G).

Ideal and Model Ligand Representations

Various files associated with chemical components are available for download from the RCSB PDB website. Coordinates of the 'ideal' and 'model' versions of the ligand are available (Figure 6). Ideal coordinates are calculated by software based on the known covalent geometry (typically using the Molecular Networks Corina or OpenEye Omega programs). These coordinates are often used in modeling applications where researchers want to minimize any deviations from ideal geometry that may be present in experimental structure determinations. Experimental coordinates are taken from the first entry in which the component was observed, and as such can represent the conformation that the ligand adopts upon binding to a macromolecule. The dictionary definition for chemical components can also be downloaded in PDBx/mmCIF and XML formats.
Ligand of Interest (LOI)
A small molecule is termed a Ligand of Interest (LOI) when it is the subject of the author’s research rather than a byproduct of the experimental conditions. For example, in entry 1o8v, the small molecule palmitic acid. (PLM) is designated as an LOI as it is bound inside the cavity of the the fatty-acid-binding protein EgFABP1 and is integral to the study.



