Molecule of the Month: Complex I
A proton-pumping protein complex performs the first step of the respiratory electron transport chain
Electron Transport
Tandem Proton Pumping

Managing Protons
Exploring the Structure
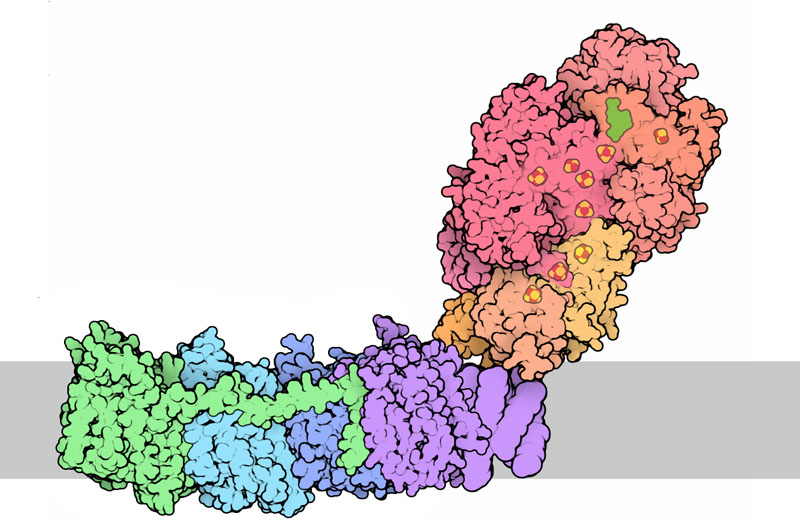
Complex I (PDB entry 3m9s)

The structure of bacterial complex I is available in PDB entry 3m9s . The structure was solved at low resolution, so not all portions of the chain were visible, and you may notice that some parts of the chain are missing. The two portions of the molecule have also been solved separately at higher resolution: you can take a look at them in PDB entries 2fug and 3rko . Click on this image for an interactive Jmol.
Topics for Further Discussion
- You can use the "overlap protein" tool at the PDB to compare the structures of three of the proton pumps: chains L, M, and N in PDB entry 3rko. The fourth proton pump is thought to be formed between several protein chains: N, K, J and A.
- The structure of complex I fills out the entire electron transport chain: you can now explore the structure of all three complexes at the PDB.
Related PDB-101 Resources
- Browse Biological Energy
- Browse Peak Performance
References
- U. Brandt (2006) Energy converting NADH:quinone oxidoreductase (complex I). Annual Review of Biochemistry 75, 69-92.
- R. G. Efremov, R. Baradaran and L. A. Sazanov (2010) The architecture of respiratory complex I. Nature 465, 441-445.
December 2011, David Goodsell
http://doi.org/10.2210/rcsb_pdb/mom_2011_12


