The Search for Drugs to Fight COVID-19
Medical researchers are testing previously-discovered antiviral drugs for effectiveness against SARS-CoV-2.
We are in the midst of a global pandemic caused by SARS-CoV-2, a newly-emerged coronavirus closely related to SARS-CoV that came and went in the early 2000s. Fortunately, the research community is working overtime to find ways to combat this new plague, but effective antiviral agents are difficult to find. Drugs need to be highly specific, so that they can cure (or at least control) disease without causing unwanted side effects. Investigational agents (would-be drugs) typically go through many phases of testing and validation before approval, to ensure that they are both safe and effective. This challenging process can be excruciatingly slow, particularly as we face a virus that is killing many thousands of people a day worldwide. One solution is to test previously-discovered drugs to see if they are effective against the new coronavirus. This approach has proven successful in other grave situations. For example, AZT was initially developed as a potential anticancer agent, but later was rapidly approved for use early in the fight against HIV. Improved variants of AZT are now allowing individuals with AIDS to live without fear of the disease or spreading the virus to others.
Many Tries
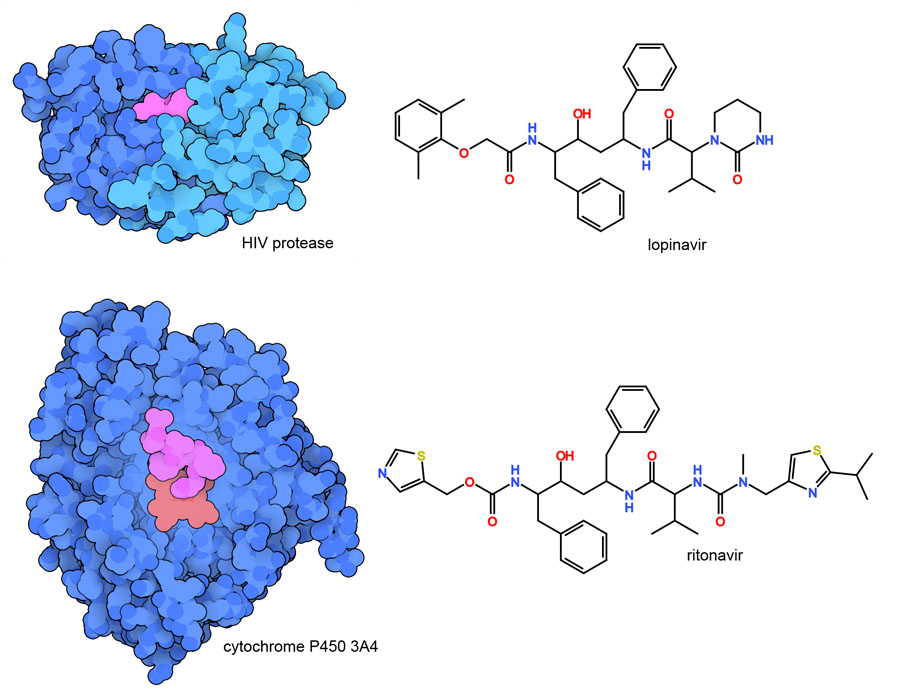
The medical community is currently trying to repurpose many approved drugs to fight the pandemic. Obvious choices include antiviral drugs that are used to treat other infectious diseases, such as the combination of lopinavir and ritonavir shown here, which are effective for treatment of HIV. Both of these compounds are designed to block HIV protease, and ritonavir also has the added advantage of blocking enzymes that metabolize drugs (PDB ID 3nxu). Unfortunately, neither HIV protease inhibitor showed success in treating COVID-19. Structures of these drugs binding to their HIV protease target (PDB ID 1mui) help us understand why this combination is not effective: HIV protease is structurally very different from the SARS-CoV-2 protease. Repurposing agents that block HIV protease was always a long shot for treatment of COVID-19, but it had to be tried. In other cases, repurposed drugs are being used successfully to fight the deadly response of the patient’s immune system to SARS-CoV-2 infection, for example, using the monoclonal antibody tocilizumab, which was developed to fight autoimmune diseases by blocking signaling by interleukin IL-6. Hundreds of clinical trials are currently under way, testing many different existing drugs for emergency use in fighting the ongoing pandemic.
Achilles Heel
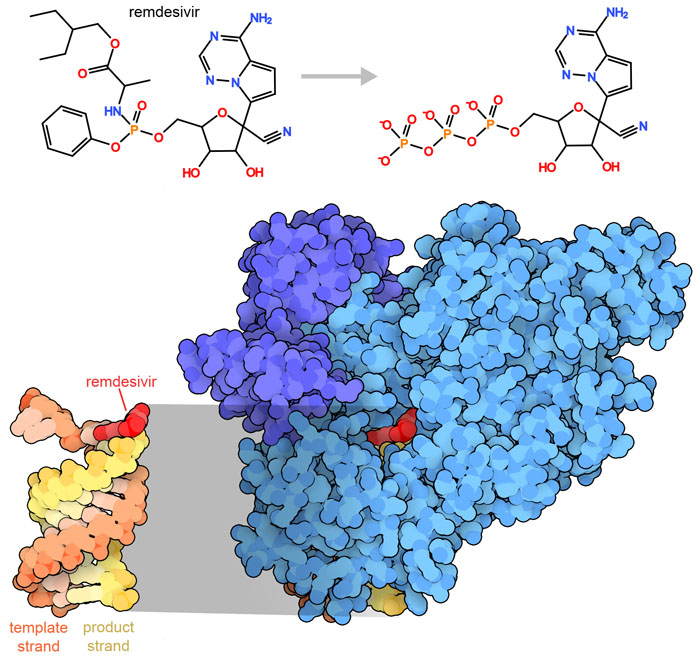
Drug hunters always look for a weak point to attack when designing and testing new antiviral agents. For coronaviruses, the RNA-dependent RNA polymerase is an attractive target, since our own cells don't have a comparable RNA polymerase. Remdesivir was initially discovered and developed to block the actions of similar enzymes found in other RNA viruses (e.g., Ebola), but it was not effective enough to be approved as an anti-Ebola drug. Fortunately, it is effective against SARS-CoV-2 in severely ill patients and was approved by the US Food and Drug Administration in October 2020 for the treatment of COVID patients. The structure shown here (PDB ID 7bv2) captures remdesivir as it’s doing its job. Remdesivir is an adenine nucleotide analog, which is a prodrug that’s converted inside the patient’s cells to its active triphosphate form. It is then added to the growing viral RNA chain, corrupting the action of the polymerase.
Building on Past Experience
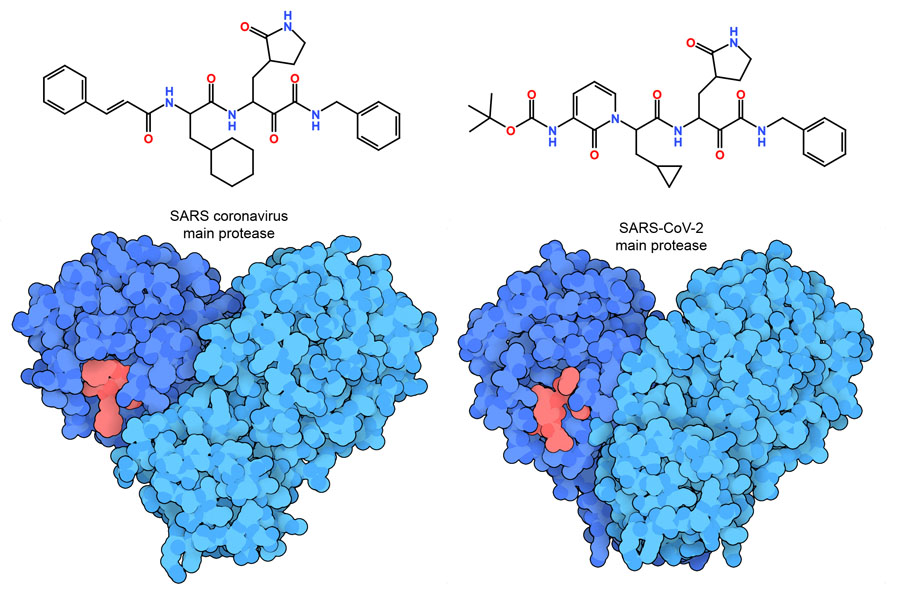
Biomedical researchers have accumulated a vast body of knowledge about viral proteins and small chemicals that inhibit them, which can be used as a foundation for discovering effective SARS-CoV-2 antivirals. For example, the ketoamide compound shown here is a potent inhibitor for the SARS coronavirus main protease (PDB ID 5n19). Researchers are now shuffling atoms around in this small chemical, in the hope of making new inhibitors effective against the SARS-CoV-2 main protease (PDB ID 6y2f). Since these two viral proteases are extremely similar, structural knowledge garnered from SARS provides an excellent head start in the fight against the new coronavirus. See the Molecule of the Month for more information on the coronavirus main protease.
More Information
- Burley, S. K. (2020) How to help the free market fight coronavirus. Nature 580: 167.
- Guy, R.K., DiPaola, R. S., Romanelli, F., Dutch, R.E. (2020) Rapid repurposing of drugs for COVID-19. Science 368, 829-830.
- Lythgoe, M. P., Middleton, P. (2020) Ongoing clinical trials for the management of the COVID-19 pandemic. Trends Pharmacol Sci 41, 363-382.
- Scavone, C., Brusco, S., Bertini, M., Sportiello, L., Rafaniello, C., Zoccoli, A.m Berrino, L., Racagni, G., Rossi, F., Capuano, A. (2020) Current pharmacological treatments for COVID-19: What’s next? Br J Pharmacol doi: 10.1111/bph.15072, online ahead of print.
Entries from the PDB Archive Discussed in this Article
- 7bv2: Yin, W., Mao, C., Luan, X., Shen, D.D., Shen, Q., Su, H., Wang, X., Zhou, F., Zhao, W., Gao, M., Chang, S., Xie, Y.C., Tian, G., Jiang, H.W., Tao, S.C., Shen, J., Jiang, Y., Jiang, H., Xu, Y., Zhang, S., Zhang, Y., Xu, H.E. (2020) Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368: 1499-1504
- 6y2f: Zhang, L., Lin, D., Sun, X., Curth, U., Drosten, C., Sauerhering, L., Becker, S., Rox, K., Hilgenfeld, R. (2020) Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved alpha-ketoamide inhibitors. Science 368: 409-412
- 3nxu: Sevrioukova, I.F., Poulos, T.L. (2010) Structure and mechanism of the complex between cytochrome P4503A4 and ritonavir. Proc Natl Acad Sci U S A 107: 18422-18427
- 1mui: Stoll, V., Qin, W., Stewart, K.D., Jakob, C., Park, C., Walter, K., Simmer, R.L., Helfrich, R., Bussiere, D., Kao, J., Kempf, D., Sham, H.L., Norbeck, D.W. (2002) X-ray Crystallographic Structure of ABT-378 (Lopinavir) Bound to HIV-1 Protease Bioorg Med Chem 10: 2803-2806



