RNA Chaperone NMB1681
July 2011

Chaperoning Duties
NMB1681 is thought to act like the similar RNA chaperone FinO. It performs several functions. First, it binds to a stem-loop structure in a small regulatory RNA, protecting it from degradation by cellular ribonuclease enzymes. Then, it pairs the small RNA with a complementary sequence in a messenger RNA, which also forms a distinctive stem-loop structure. The complex of the two RNA molecules hides the Shine-Delgarno sequence and thus reduces production of proteins from the message.Structured and Unstructured
Both NMB1681 and FinO are composed of two parts, which perform different aspects of their function. The core of NMB1681, recently solved by MCSG researchers and shown here from PDB entry 3mw6, performs the first task, recognizing the stem-loop structure and protecting it. The rest of the protein forms an unstructured tail that is essential for the chaperoning task of bringing the two RNA molecules together. This type of unstructured protein is actually quite common in RNA chaperones, and may play a role in the process of folding and unfolding of RNA secondary structure like the stem-loop structures.Complementary Charge
The NMB1681 has a distinctive shape and chemical composition. It is saddle-shaped, with a dense collection of arginine, lysine, and histidine amino acids (shown above in blue) on the curve of the saddle. These amino acids normally carry a positive charge, which is perfect for stabilizing the many negatively-charged phosphates on the RNA strands. Serendipitously, the crystal lattice in this structure solution includes six independent copies of the protein, so the structure also gives a glimpse at the essential flexibility of the protein.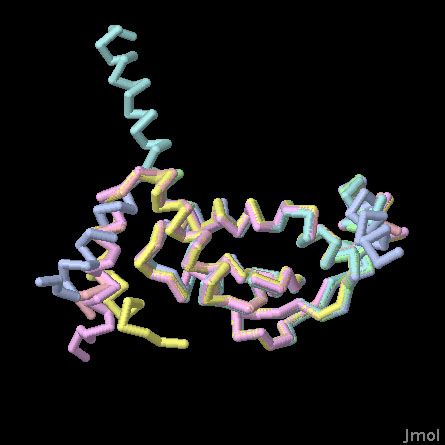
RNA Chaperone NMB1681 (PDB entry 3mw6)
The crystal structure was solved using a fragment of the NMB1681 that includes the RNA-binding core and a short segment of the flexible tail. In this Jmol, the cores of the six independent structures are overlapped, so you can see some of the range of motion of the flexible tail. Use the buttons to turn the structures on and off, and to look at a spacefilling representation that shows the positively-charged amino acids.
References
- Chaulk, S. et al. N. meningitidis 1681 is a member of the FinO family of RNA chaperones. RNA Biology 7, 812-819 (2010).
- Ghetu, A. F., Gubbins, M. J., Frost, L. S. & Glover, J. N. M. Crystal structure of the bacterial conjunction repressor FinO. Nature Structural Biology 7, 565-569 (2000).



