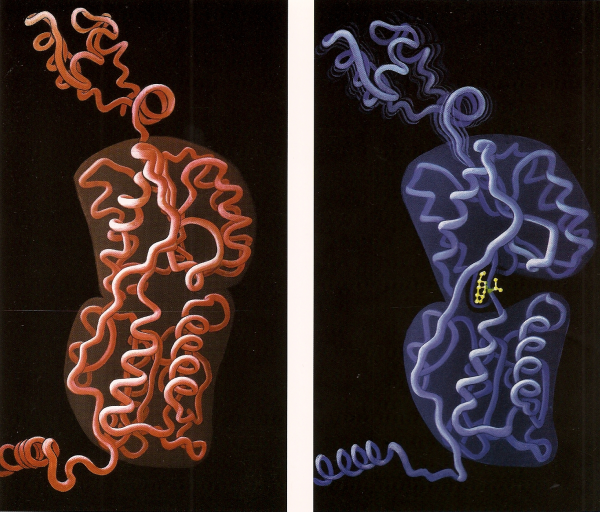
Induced Lac Repressor
Unknown Year, 15 ½”, 10”

This was Geis’ last completed painting. It depicts the structure of a lac repressor monomer, including the tetramerization alpha helix at the bottom, the middle C-domain and N-Domain, and the headpiece domain at the top which binds the lac operator DNA. On the left is the repressed state, which has a high affinity to DNA. On the right is the induced state in which IPTG (isopropyl beta-D-1-thiogalactopyranoside), a artificial inducer, causes a conformational change and reduces the affinity of the repressor for the lac operator, turning on a gene which increases lactose metabolism in bacteria cells. Geis uses a clean tubular style, making it easier to see the conformational change. IPTG causes the gap between the C-subdomain and the N-subdomain to widen. This destabilizes the bonds between the headpieces and the DNA target, shown by multiple shadows emanating from the headpiece residues.
Used with permission from the Howard Hughes Medical Institute (www.hhmi.org). All rights reserved.

Related PDB Entry: 2P9H
Experimental Structure Citation
Daber, R., Stayrook, S., Rosenberg, A., & Lewis, M. (2007). Structural Analysis of Lac Repressor Bound to Allosteric Effectors. Journal of Molecular Biology,370(4), 609–619.
About Lac Repressor
The lac repressor is a protein that can bind to certain DNA sites to prevent binding of RNA polymerase. This can therefore prevent transcription of genes for lactose metabolism, until further action from other proteins. The tetramer structure forms a V-shape, with the headpieces attaching to DNA and possibly twisting the DNA into a loop.
Text References
Gilbert, W., & Müller-Hill, B. (1967). The lac operator is DNA. Proceedings of the National Academy of Sciences of the United States of America, 58(6), 2415–2421.Lewis, M. (2011). A tale of two repressors – a historical perspective. Journal of Molecular Biology, 409(1), 14–27. http://doi.org/10.1016/j.jmb.2011.02.023
Bell, C. & Lewis, M. (2001). The Lac repressor: A second generation of structural and functional studies. Current Opinion in Structural Biology 11(1):19-25.