Molecule of the Month: Beta-galactosidase
Beta-galactosidase is a powerful tool for genetic engineering of bacteria

Discovering Operons
Dyes and Complementation
Engineering Bacteria
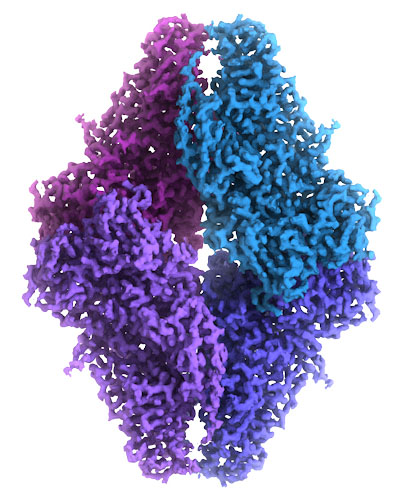
Atomic EM
Exploring the Structure

The structure of beta-galactosidase revealed how this process of complementation works. PDB entry 1jz7 is shown here, which includes a molecule of galactose in the active site, shown here in white and pink spheres. The small peptide used for complementation, shown here in green and magenta, passes through a tunnel in the protein, stabilizing the active, tetrameric form of the enzyme. To explore this structure in more detail, click on the image for an interactive JSmol.
Topics for Further Discussion
- The mechanism of the cleavage reaction has been studied with complexes of the enzyme with a variety of reaction intermediates. Try going to entry 4v44 and use the “Sequence Similarity” tab to find these structures.
- The large beta-galactosidase enzyme may have evolved from smaller enzymes that perform similar reactions. Try searching for “beta-galactosidase” to see some.
Related PDB-101 Resources
- Browse Integrative/Hybrid Methods
- Browse Enzymes
- Browse Biological Energy
- Browse Recombinant DNA
References
- 5a1a: A. Bartesaghi, A. Merk, S. Banerjee, D. Matthies, X. Wu, J. L. S. Milne & S. Subramaniam (2015) 2.2 A resolution cryo-EM structure of beta-galactosidase in complex with a cell-permeant inhibitor. Science 348, 1147-1151.
- D. H. Juers, B. W. Matthews & R. E. Huber (2012) LacZ beta-galactosidase: structure and function of an enzyme of historical and molecular biological importance. Protein Science 21, 1792-1807.
- 1jz7, 1jz8: D. H. Juers, T. D. Heightman, A. Vasella, J. D. McCarter, L. Mackenzie, S. G. Withers & B. M. Matthews (2001) A structural view of the action of Escherichia coli (lacZ) beta-galactosidase. Biochemistry 40, 14781-14794.
June 2016, David Goodsell
http://doi.org/10.2210/rcsb_pdb/mom_2016_6


