2020 Video Challenge for High School StudentsMolecular Mechanisms
|
2020 Video Challenge
Winners | Entries | Judges | Overview | Learning Resources
Note: Videos are intended for communication and not rigorous scientific review.
Congratulations to the 2020 Winners:
2020 Challenge Details
In this challenge, we ask you to tell a story that communicates:
-
Opioid receptor agonists: similarities and differences between endogenous opioids and opioid drugs and their interaction with opioid receptors
-
GPCR signaling: introduce one molecular mechanism of pain signal modulation induced by the activation of the G-protein by opioids. This might include:
- Preventing of neurotransmitter release
- Modification of the action of potassium channel
- Modification of the action of adenylyl cyclase
Opioid receptor antagonists: how Naloxone works to prevent death from opioid overdose
Opioid crisis: ways to fight opioid crisis and prevent opioid abuse
2020 Dates
The video submission opened on January 14, 2020 and concluded on April 28, 2020. Award winners were announced at rcsb.org on May 19, 2020.
2020 Learning Resources
I. Neuronal Signaling
There are 2 important processes involved in carrying out the neuronal signal: intracellular signaling (within the cell) and intercellular signaling (between the cells). Understanding of these two processes is essential for comprehending how opioid signaling modulates the neuronal transmission pathways. Below you will find a brief overview of both processes. More information about these topics can be found here and here.
Intracellular signaling
Figure 1

Schematic drawing of a neuronal cell with a ~50 nm section of the axon highlighted. The enlargement shows the cell in the resting state highlighting the sodium (green) and potassium (magenta) ion gradients on the extracellular and intracellular side of the membrane. Proteins essential for intracellular signal transmission are shown: A: voltage gated sodium channel (PDB structure 5vb8), B: voltage gated potassium channel (PDB structure 5k7l), and C: sodium-potassium pump (PDB structure 2zxe).
The intracellular signaling is achieved by a progressive travel of positive charge - called action potential - along the neuronal membrane. The signaling is regulated by electrically charged particles: potassium and sodium ions, with sodium concentration higher on the outside and potassium higher on the inside of the cell membrane (Figure 1). When the neuron is at rest (resting potential) the electrical potential on the inside is lower by about 70 mV relative to the outside and the membrane is said to be polarized.
The action potential is a sudden spike in the electrical potential on the inside of the neuron caused by a sudden influx of positively charged ions. Action potentials are generated/regulated by voltage gated ion channels which contain voltage sensing domains that react to electrical stimulus by opening up and allowing ions to enter or exit the cell. Once their threshold has been reached they become inactivated and can’t conduct any signal for a while.
The voltage gated sodium channel (Figure 1A) is essential in facilitating the travel of the action potential down the axon. Upon its activation sodium enters the cell raising the inside voltage. The channels trigger one another and allow the sodium to depolarize the membrane along the axon.
The repolarization of the membrane is facilitated by the voltage gated potassium channels (Figure 1B). These channels open at a higher voltage and allow potassium ions to exit the cell, lowering the potential. The sodium-potassium pump (Figure 1C) restores the sodium and potassium gradients on the inside and the outside.
Intercellular signaling
Figure 2
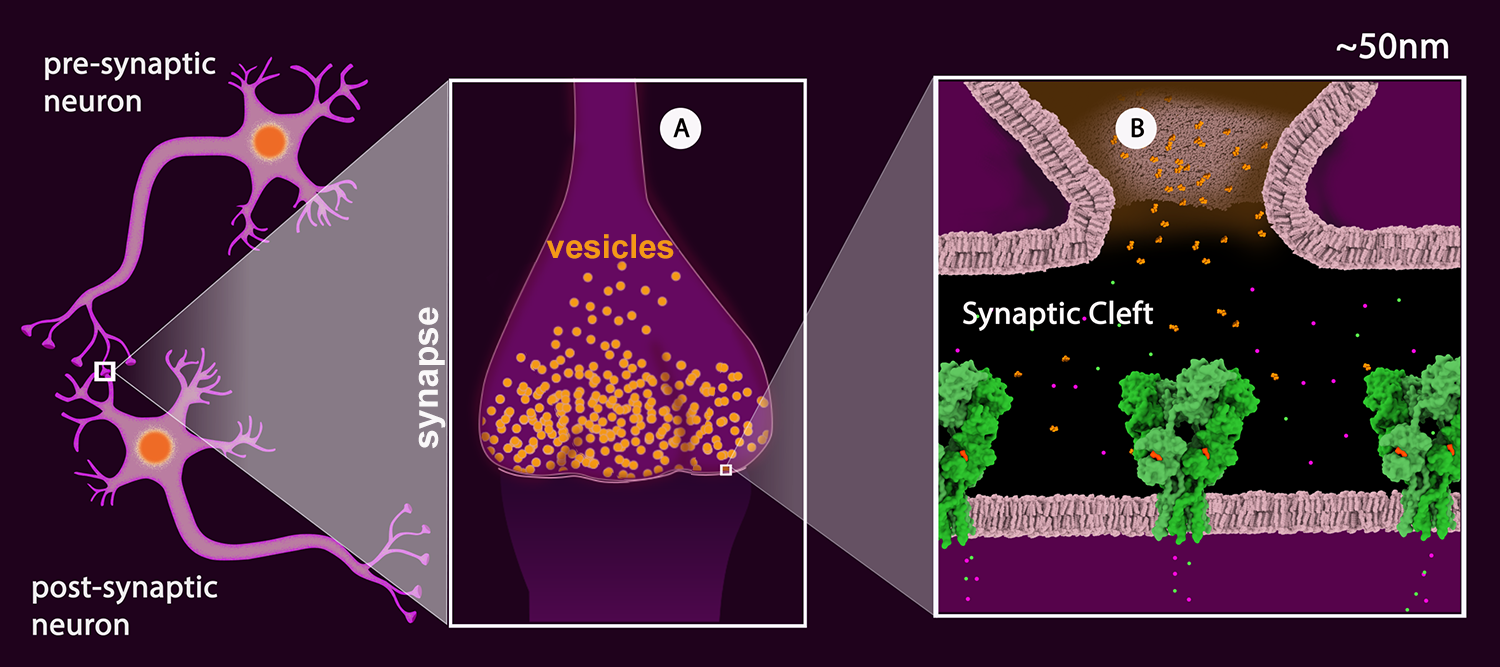
Schematic drawing of a neuronal junction with the synaptic connection highlighted. A: The synapse with synaptic vesicles (orange). B: View of the synaptic cleft showing one vesicle merged with the pre-synaptic membrane and the neurotransmitter (glutamate - orange) released into the cleft. The neurotransmitter receptors are shown in green. (AMPA Receptor, PDB Structure 3kg2). The receptor shown is a ligand-gated ion channel permeable to sodium (green) and potassium (magenta).
Once the action potential reaches the synapse of the pre-synaptic neuron, the signal is passed onto the post-synaptic neuron (Figure 2) by the release of neurotransmitters. Neurotransmitters are chemical substances that are stored in synaptic vesicles (Figure 2A). These vesicles fuse with the pre-synaptic neuron membrane releasing neurotransmitters into the presynaptic cleft (Figure 2B).
The post-synaptic membrane contains neurotransmitter receptors. These receptors can be ionotropic or metabotropic. Ionotropic receptors, also called ligand gated ion channels, contain an ion channel that opens upon neurotransmitter binding allowing the ions to flow through the channel (e.g. AMPA Receptor, Figure 2B). Metabotropic receptors, also called G-protein coupled receptors or GPCRs, activate signal transduction upon neurotransmitter binding (e.g. Serotonin Receptor).
Depending on the type of neurotransmitter stored by the pre-synaptic neuron and type of receptors present on the post-synaptic neuron, the synapse can be excitatory or inhibitory. In the excitatory synapse, the excitatory neurotransmitter receptor allowing the positively charged ions to enter the neuron, generating the action potential (e.g. AMPA Receptor, Figure 2B). In the inhibitory synapses, the neurotransmitter stops the signal upon binding to the receptor (e.g. Glutamate-gated Chloride Receptors), usually by allowing negatively charged ions to enter the cell to hyperpolarize the membrane (increase the difference in voltage on the inside relative to the outside beyond the resting potential).
The release of neurotransmitters is dependent on the presence of calcium ions inside the pre-synaptic cell, as they activate proteins that enable the fusion of the neurotransmitter vesicle with the pre-synaptic membrane. The concentration of calcium is high on the outside and very low on the inside. The voltage gated calcium channels on the pre-synaptic membrane respond to the action potential by opening up and allowing calcium to enter the cell.
II. Endogenous and exogenous opioids and opioid receptors
Endogenous opioids (made by our bodies) are short chains of amino acids known as endorphins, enkephalins, and dynorphins. These neurotransmitters are produced and stored in large core vesicles inside certain neurons in the central nervous system, and pituitary and adrenal glands. They are not typically released into the synaptic cleft like GABA or glutamate, but rather into the pre- and post-synaptic areas where they can modulate the primary communication pathway, altering our response to pain or stress.
Exogenous opioids also called opioid drugs can be derived from plants (e.g. morphine is derived from opium poppies) or produced synthetically (e.g. DAMGO).
Figure 3
Examples of endogenous and exogenous opioids. Left: endogenous opioid peptide, met-enkephalin from PDB entry 2LWC with backbone carbons highlighted in black. Center: plant-derived opioid morphine. Right: synthetic opioid peptide, DAMGO with peptide backbone highlighted in black (chain D from entry 6DDF).
Both, endogenous and exogenous opioids interact with opioid receptors that are present on the pre- and post-synaptic membranes in central and peripheral nervous systems.
All opioid receptors are part of a larger family of proteins called G-protein coupled receptors (GPCRs). All GPCRs have structural features that have been preserved through the course of evolution. With the extracellular N-terminus, the protein chain folds to form a bundle of seven transmembrane alpha helices connected by 3 intracellular and 3 extracellular loops with the C-terminus reaching inside the cell. On the extracellular side, the helices form a cavity where ligands (e.g. endorphins, morphine) bind. On the intracellular side, the receptor is coupled to G-protein. Several types of opioid receptor types have been found. The three receptors with high affinity for morphine-like compounds are listed below.
Table 1: Types of opioid receptors
| Type of receptor and example PDB Structure | Agonists Examples (bold indicates the highest potency) |
Visualization hints and resources |
Learning Resources |
Mu-opioid receptor 4dkl |
Endogenous: Exogenous: |
Chimera session (preview below)b The Chimera session preview for the PDB structure 4dkl shows the mu-opioid receptor in dark cyan (chain A). The N- and C- termini are colored in blue and red respectively. The receptor has an experimental antagonist beta-funaltrexamine bound covalently to Lys 233 in the active site (purple – ligand BF0). An engineered version of the receptor was used to solve this structure, with lysozyme (white) inserted in the loop between helices 5 and 6; it is not normally a part of the structure. The opioid receptors are thought to exist as dimers. To see the second unit, go to Favorites > Model Panel and choose Biological Unit from the menu on the right. | Molecule of the Month on Opioid Receptors 2019 calendar What is a Protein? page for November G Protein-Coupled Receptor (GPCR) Paper Model
|
Kappa-opioid receptor 4djh |
Endogenous: Exogenous: |
Chimera session (preview below)b The Chimera session preview shows the kappa-opioid receptor structure 4djh. The opioid receptors are thought to function as dimers. One subunit (chain A) is colored dark cyan, the second (chain B) is colored in cyan. The N- and C- termini are colored in blue and red respectively. The receptor has an antagonist JDTic bound in the active site (purple, ligand JDC). An engineered version of the receptor was used to solve this structure, with lysozyme (white) inserted in the loop between helices 5 and 6; it is not normally a part of the structure. |
|
Delta-opioid receptor 4ej4 |
Endogenous: Exogenous: |
Chimera session (preview below)b The Chimera session preview shows the delta-opioid receptor structure 4ej4 in dark cyan (chain A). The N- and C- termini are colored in blue and red respectively. The receptor has a highly selective delta opioid receptor antagonist naltrindole bound in the active site (purple – ligand EJ4). An engineered version of the receptor was used to solve this structure, with lysozyme (white) inserted in the loop between helices 5 and 6; it is not normally a part of the structure. |
III. Molecular Mechanisms of opioid action: GPCR Signaling
On the intracellular side, opioid receptors are coupled to G-proteins. G-proteins are trimeric complexes consisting of alpha (Gα), beta (Gβ), and gamma (Gγ) subunits. When inactive, they have GDP bound to the alpha subunit. Opioid binding induces a conformational change in the receptor that leads to the activation of the G-protein. Upon activation the GDP is released and GTP binds instead and the complex splits into 2 parts: Gα and Gβγ. By integrating with other proteins and channels, they modulate the neuronal pathways, altering our perception of pain or stress.
Within the Gα subunit, there are a few classes, for example the stimulatory Gα (Gαs) or the inhibitory Gα (Gαi). As the names suggest, upon binding to other proteins within the signaling pathways they stimulate the protein function or inhibit it. All three types of opioid receptors listed in Table 1 interact with Gαi.
Examples of neuronal signal modulation mechanisms by opioids are listed below:
-
Preventing neurotransmitter release
If the receptor is activated on the pre-synaptic neuron, the Gβγ can bind to voltage gated calcium channels blocking the voltage sensing domains and preventing calcium to enter the cell. As the neurotransmitter release is dependent on intracellular calcium, this can stop the signal from being passed on from one neuron to another.
-
Modification of the action of potassium channel
The Gβγ regulate the activity of some potassium channels. When they bind to them, the channels remain open allowing potassium to exit the cell. This leads to hyperpolarizing of the membrane (raising the difference between the inside and outside of the neuron to more than 70 mV). This results in dissipation of the action potential.
-
Decreasing cyclic AMP (cAMP) production
Neurotransmitters are often termed the first messengers. The signal generated by them is amplified by intracellular molecules called second messengers. An example of such second messenger is cAMP. cAMP is synthesized from ATP by the enzyme adenylyl cyclase, once the enzyme is activated. cAMP can bind to some channels directly and activate others indirectly via cAMP dependent protein kinases. Both activities augment the ion transport and thus strengthen the neuronal signal. The Gαi can bind to adenylyl cyclase inactivating the enzyme and preventing the production of cAMP.
Table 2: Example PDB entries for G-protein signaling and molecular mechanisms of neuronal signal modulation by opioids
|
Type of the protein and example PDB Structure(s) |
Visualization hints and resources |
Learning resources |
Mu-receptor with inhibitory G-protein 6dde 6crk |
6dde 3D view from NGLa Chimera session (preview below)b The Chimera session preview shows the mu-receptor with G-protein from the PDB entry 6dde. The opioid receptor is shown in dark cyan (chain R). An agonist DAMGO (chain D, purple) is bound in the active site. The G-protein subunits are shown underneath the receptor. Gαi is shown is salmon (chain A), Gβ is shown in light green (chain B), Gγ is shown id darker green (chain C). The PDB structure also contains an antibody fragment (chain E) that was added to the protein for crystallization, however it is not a part of biological molecule and it is hidden in this session. The structure 6dde has an incomplete Gα subunit. The PDB structure 6crk shows a full-length G-protein. The Gα comprises RAS domain (where GTP/GDP binds) and alpha-helical domain (colored in peach). The GDP is shown in its binding site. Both structures are aligned in the Chimera session. Use Favorites > Model Panel with the Shown checkboxes to toggle between these 2 structures. |
Molecule of the Month on G-Proteins Flyer on G-Protein Coupled Receptors |
Voltage Gated Calcium Channel 6jp5 |
Chimera session (preview below)b The Chimera session preview shows the structure 6jp5. The voltage gated calcium channels comprise multiple subunits: alpha-1, alpha-2, beta-1, delta-1, and gamma-1. The alpha-1 subunit of the channel forms the pore and is colored in light purple (chain A). Two calcium ions (green) are shown inside the pore. Highlighted in orange are the voltage gating domains with the positively charged side chains that react to voltage increases shown in ball and stick representation. The beta-1 subunit (purple, chain B and C) regulates the activity of the pore forming subunit. The alpha-2/delta-1 (chain F, light blue) and gamma-1 (chain E, pink) subunits regulate the channel inactivation. There is no PDB structure at present capturing the interaction of Gβγ complex with a voltage gated calcium channel. Research shows that the Gβγ complex interacts with the alpha-1 subunit (learn more).
|
|
Potassium channel in complex with G-beta-gamma subunits 4kfm |
Chimera session (preview below)b The Chimera session preview for the structure 4kfm shows the Gβγ subunits (light green, dark green – chains B and G, respectively) bound to a potassium channel (chain A, purple). The channel consists of 4 identical subunits (chain A x 4). Four potassium ions are shown inside the channel in magenta. In addition to Gβγ subunits bound, special membrane phospholipids (ligand PIO, orange) are required for the channel regulation (learn more). |
|
Adenylyl cyclase bound to Gαs 6r3q |
Chimera session (preview below)b The Chimera session preview for the structure 6r3q shows the adenylyl cyclase (blue) with the Gαs (salmon) bound to it. Both Gαs and Gαi can bind to adenylyl cyclase, stimulating or inhibiting the production of cAMP, although only Gαi binding would be regulated by opioids. Both types use the same binding site, and crystal structures suggest that they are structurally similar, however, some molecular dynamics experiments show that they use different mechanisms in interaction with the adenylyl cyclase (learn more). At present there is no PDB structure capturing the interaction of adenylyl cyclase with Gαi. |
IV. Opioid antagonists
While the endogenous opioids are regulated by intricate systems in our bodies, the exogenous opioids enter different systems in our bodies and bind to opioid receptors indiscriminately, causing side effects. This is dangerous when opioids are not used as prescribed by a medical professional.
For example, in opioid overdose the drugs can shut down the signaling pathway between the brain and the respiratory system that ensures that we are breathing constantly. This can lead to death. If such case occurs, the drug naloxone can be administered to save a person’s life. Naloxone is similar in structure to opioids and binds in a similar location on the receptor but doesn't activate the G-protein. As a result, it makes the receptors unavailable for opioids, keeping the communication pathways between the brain and the respiratory system functional.
At present there is no PDB structure of opioid receptor with naloxone bound. However, all receptors shown in Table 1 have antagonists bound to them. You can compare their binding mode with e.g. the PDB structure 6dde which contains an agonist in the binding site.
Visualization Resources
- The user guide for NGL can be found here. The NGL is the default 3D viewer accessible from each structure summary page, from the tab “3D View”.
- You can download and open these sessions using UCSF Chimera.
Use the tutorials available here and here to edit the sessions, create animations or save pictures.
References
- Christoph Stein (2016) Opioid Receptors. Annual Review of Medicine. Vol. 67:433-451 (Volume publication date January 2016). https://doi.org/10.1146/annurev-med-062613-093100














