MurA
UDP-N-acetylglucosamine enolpyruvyl transferase (MurA) is a transferase enzyme, found in the cytoplasmic region of a bacterium, where it plays an essential role in the synthesis of the bacterial cell wall. It is currently the target of only one US FDA approved antibacterial drug: fosfomycin.
Function
MurA, which may also be known as MurZ in Escherichia coli (E. coli) (Jia et al., 2017), catalyzes the first committed step in the peptidoglycan biosynthesis pathway. The enzyme is responsible for transferring the enolpyruvyl moiety from phosphoenolpyruvate (PEP) to UDP-N-acetylglucosamine (UDP-GlcNAc) to produce enolpyruvyl UDP-GlcNAc.
There are only two known enolpyruvyl transferases, one of which is the MurA enzyme. The other enolpyruvyl transferase is 5-enolpyruvyl-shikimate-3-phosphate (EPSP) synthase, which is an enzyme that is involved in the biosynthesis of aromatic compounds in plants and microorganisms, such as bacteria and fungi (Skarzynski et al., 1996). Because both of the pathways that MurA and EPSP synthase are involved in are essential for the survival of microbes and are not found in mammals, enolpyruvyl transferases are seen as attractive targets of novel antimicrobial agents (Eschenburg et al., 2003).
Structure
Although many bacteria possess the enolpyruvyl transferase enzyme to initiate synthesis of the bacterial cell wall, not all MurA enzymes are 100% identical. The enzyme may have the same function in different bacterial species but may have additional residues or loops that cause slight structural discrepancies among MurA homologs. For instance, the polypeptide chains of MurA homologs could be of different lengths or the catalytic residues may be in slightly different locations. However, as long as the interactions within the active site remain the same for the most part, the homologs will all be able to carry out their assigned functions.
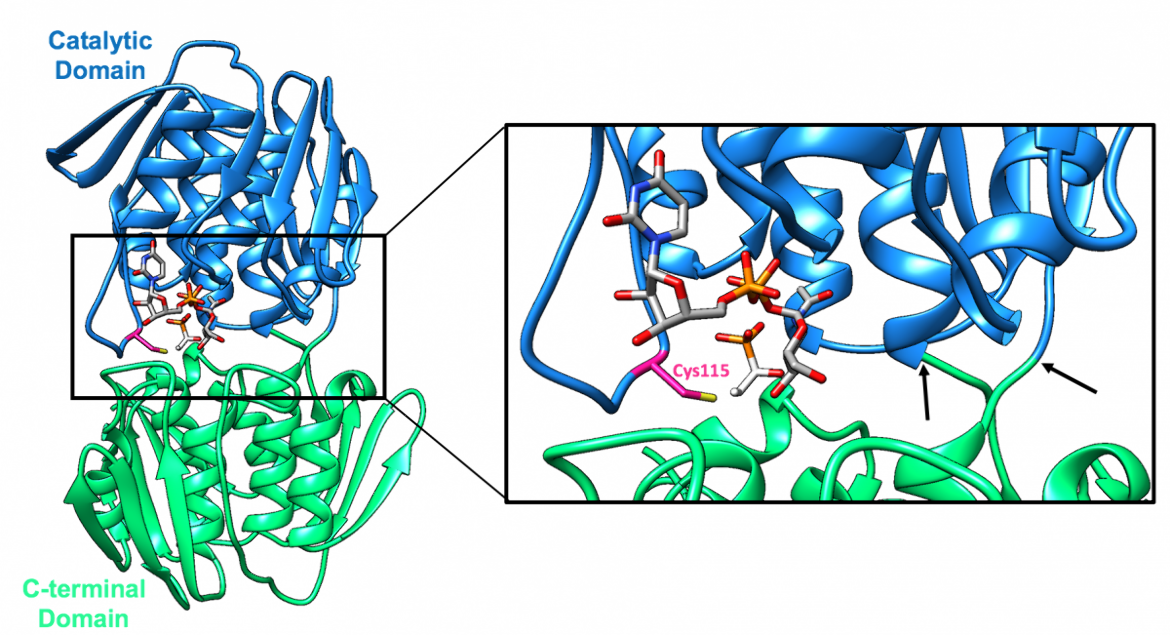
In the case of E. coli, the MurA polypeptide chain is 419 residues in length (Skarzynski et al., 1996). The single chain folds into two similar globular domains (Eschenburg et al., 2004) that are connected by a double-stranded linker, which can be seen in Figure 1 (Skarzynski et al., 1996). The refined structure, which was determined using X-ray crystallography, includes 419 residues and is in complex with UDP-GlcNAc and fosfomycin (Skarzynski et al., 1996).
The two globular domains of MurA include:
1. The catalytic domain (residues 22-229, colored blue), which contains the catalytic cysteine residue (Cys115 in E. coli and Enterobacter cloacae (E. cloacae))
2. The C-terminal domain (residues 1-21 & 230-419, colored green)
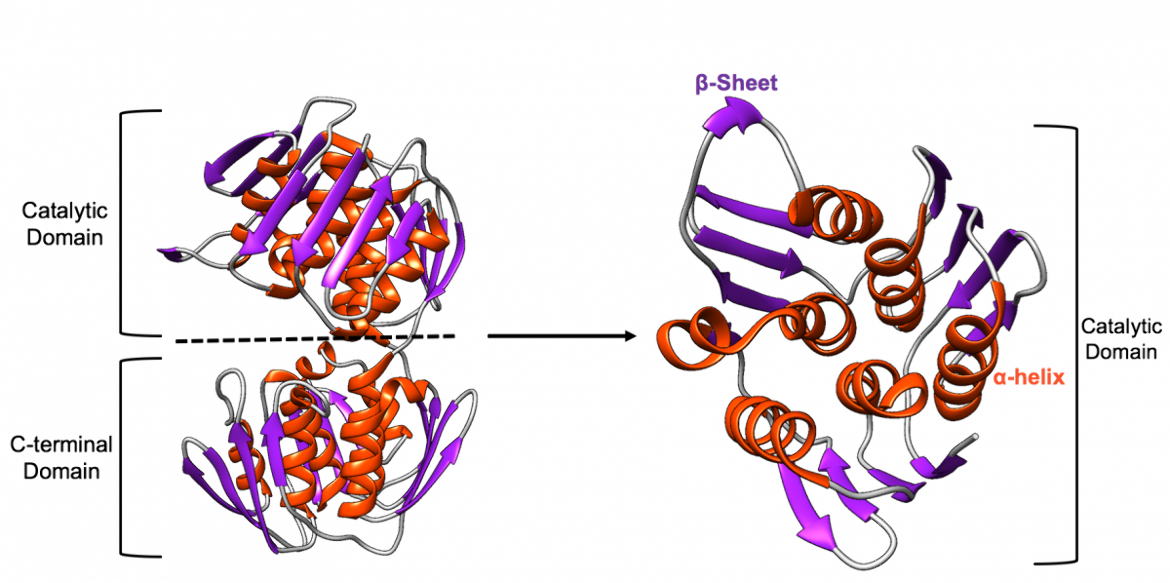
The overall fold of each domain is very similar as there are three parallel internal α (pronounced alpha)-helices that are surrounded by three additional α-helices and three four-stranded β (pronounced beta)-sheets, that are exposed to solvent, as shown in Figure 2 (Skarzynski et al., 1996).
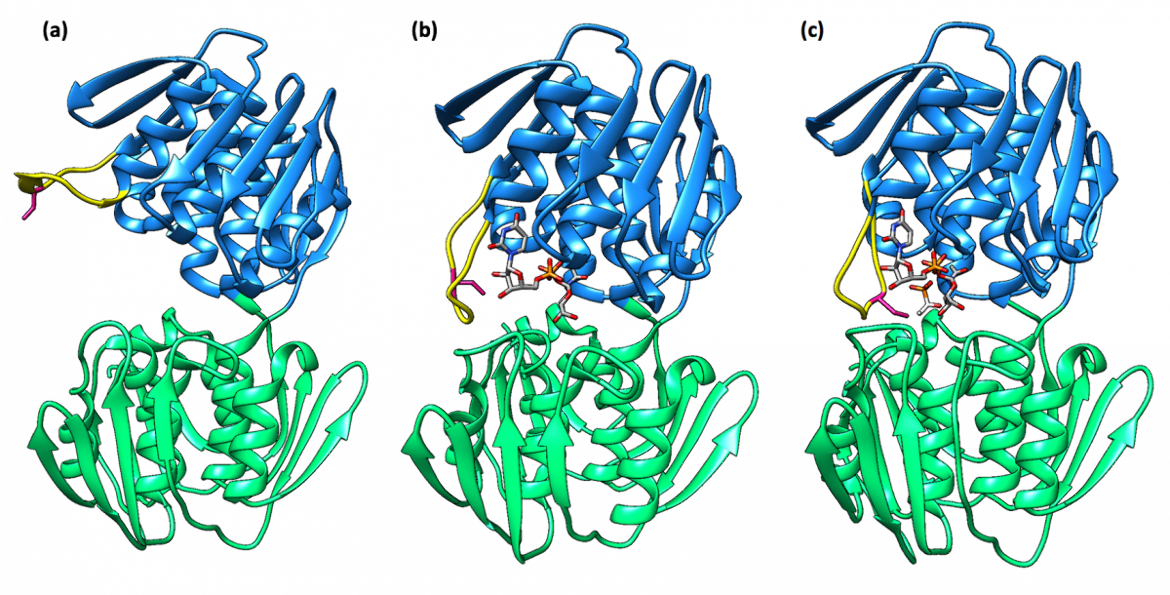
There are two distinct conformations of MurA that have been reported: an “open” conformation in the absence of substrates and a “closed” conformation in the presence of UDP-GlcNAc and fosfomycin (Samland et al., 1999). This reveals the fact that the enzyme follows an induced-fit mechanism in which the structure of MurA changes to adopt a tighter conformation as a result of substrate binding (Schönbrunn et al., 1999).
A loop region, which contains a catalytic cysteine residue (Cys115 in the case of E. coli and E. cloacae), moves as the structure undergoes conformational changes upon binding of UDP-GlcNAc (Samland et al., 1999). To be more specific, the loop is originally exposed to solvent in the ligand-free MurA structure; however, as it moves toward the UDP-GlcNAc binding site, during the transition to the closed conformation, the loop forms a tight lid around the interdomain section (Schönbrunn et al., 1999). The active site is thus created by induced fit upon binding of UDP-GlcNAc, which brings the catalytic cysteine residue into place for reaction, as depicted in Figure 3 (Skarzynski et al., 1996).
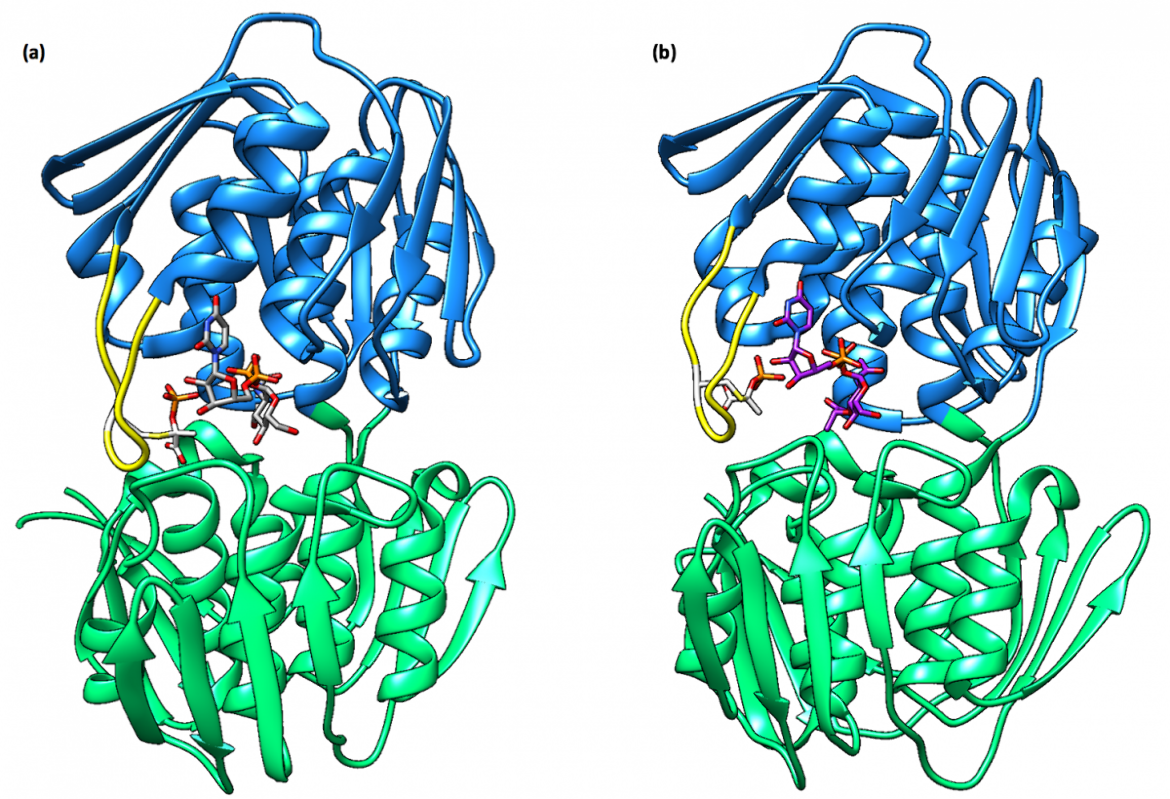
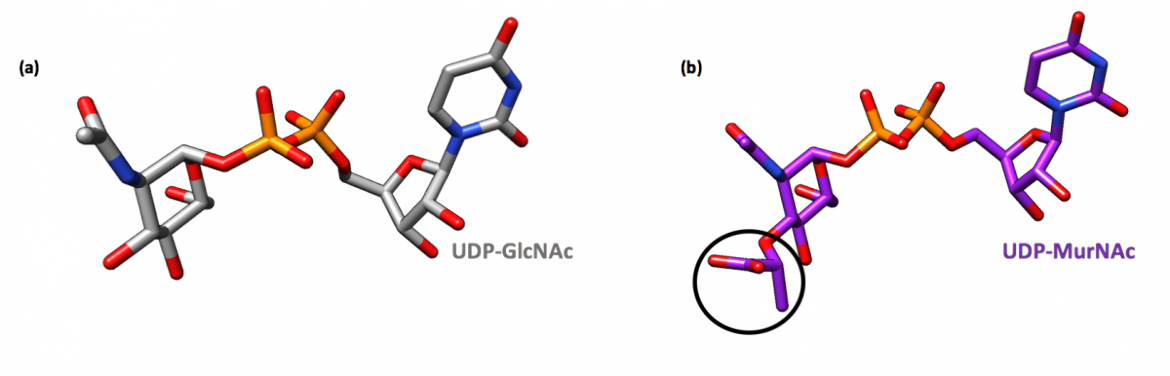
Previously, it was thought that the enzyme predominantly existed in complex with UDP-GlcNAc; however, more recent findings indicate that MurA from various different bacterial species exists largely in a dormant state bound to UDP-N-acetylmuramic acid (UDP-MurNAc) and PEP (Zhu et al., 2012). UDP-MurNAc is not only the product of the second reaction in the peptidoglycan biosynthesis pathway but also serves as a feedback inhibitor of MurA, which allows for the regulation of the peptidoglycan biosynthesis pathway (Mizyed et al., 2005). The dormant complex results from low cellular levels of UDP-GlcNAc and a simultaneous accumulation of UDP-MurNAc (Zhu et al., 2012).
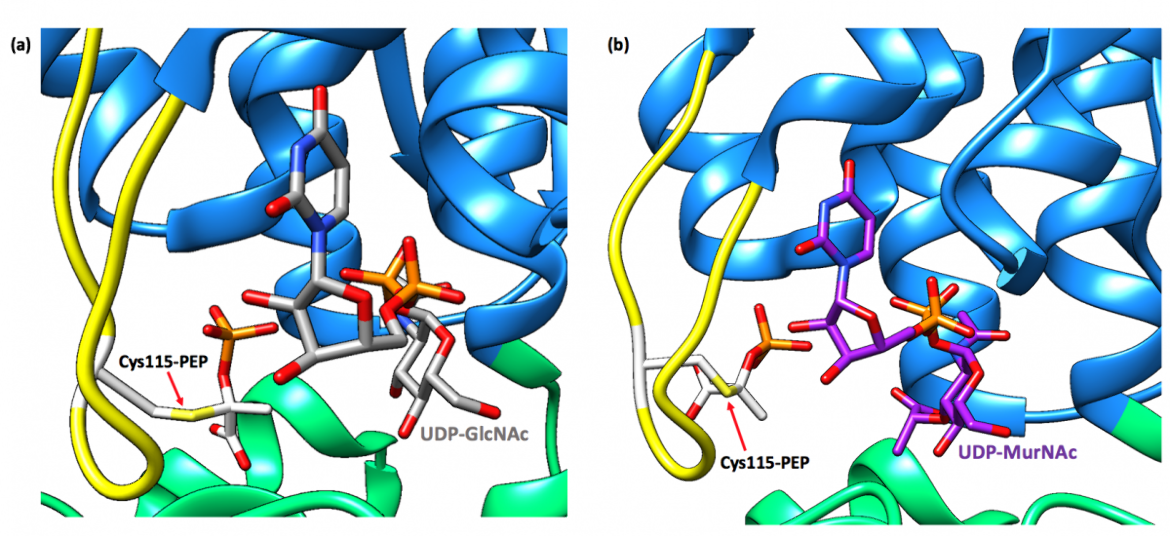
When bound to the dormant enzyme, UDP-MurNAc occupies the binding site of UDP-GlcNAc, which is seen in Figure 4b, and PEP forms a covalent adduct with the catalytic cysteine residue (Cys115 in E. coli and E. cloacae). This phospholactoyl adduct, shown in Figure 5b, is then able to form several electrostatic interactions with surrounding residues, which tightens the complex; therefore, MurA exists predominantly in a tight, resilient complex that can then only be unlocked by incubation with UDP-GlcNAc or inorganic phosphate (Zhu et al., 2012).
Once UDP-GlcNAc displaces UDP-MurNAc, PEP is transferred from the catalytic cysteine residue to UDP-GlcNAc to produce enolpyruvyl UDP-GlcNAc, all of which occurs in a single-turnover reaction (Zhu et al., 2012). These findings show that in addition to tightening the complex, the phospholactoyl adduct also activates the PEP molecule for future catalysis (Zhu et al., 2012).
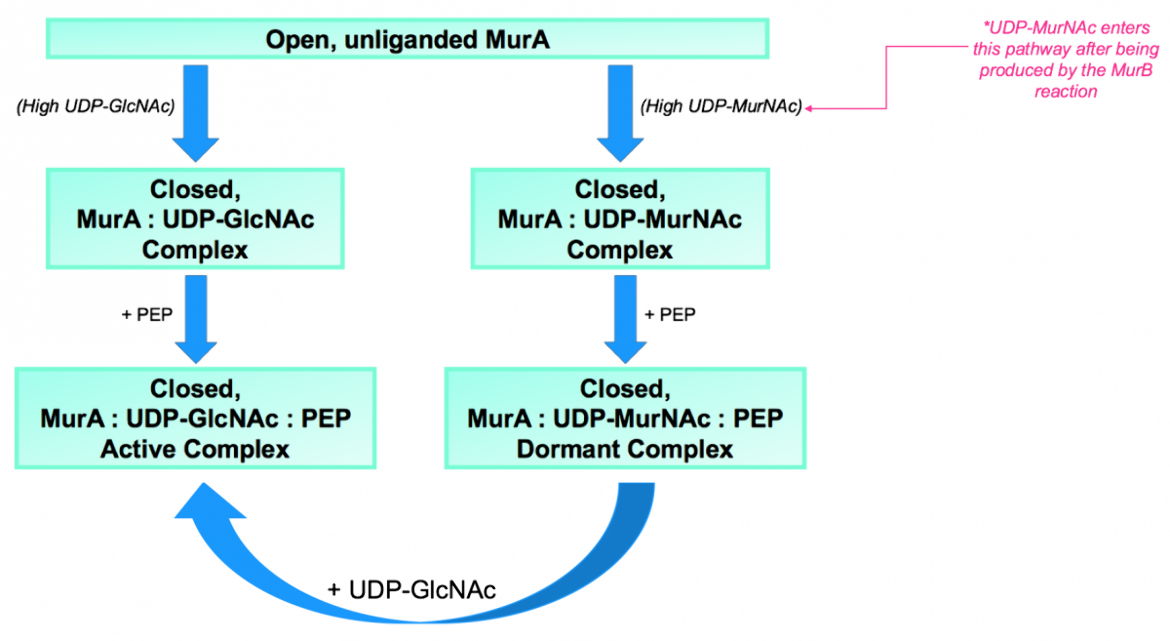
|
| Figure 7. (a) Summary of the conformations and transition states that MurA can be found in. Source: adapted from (Zhu et al., 2012). |
Active Site
As was previously noted, the catalytic site is found in the cleft between the two globular domains, where substrates and potential inhibitors will bind. Within the active site, certain direct interdomain interactions around the catalytic pocket not only stabilize the closed conformation of the substrate-bound enzyme but also control access to the active site during different stages of the reaction (Skarzynski et al., 1996). These interdomain interactions can be seen in Figure 8.
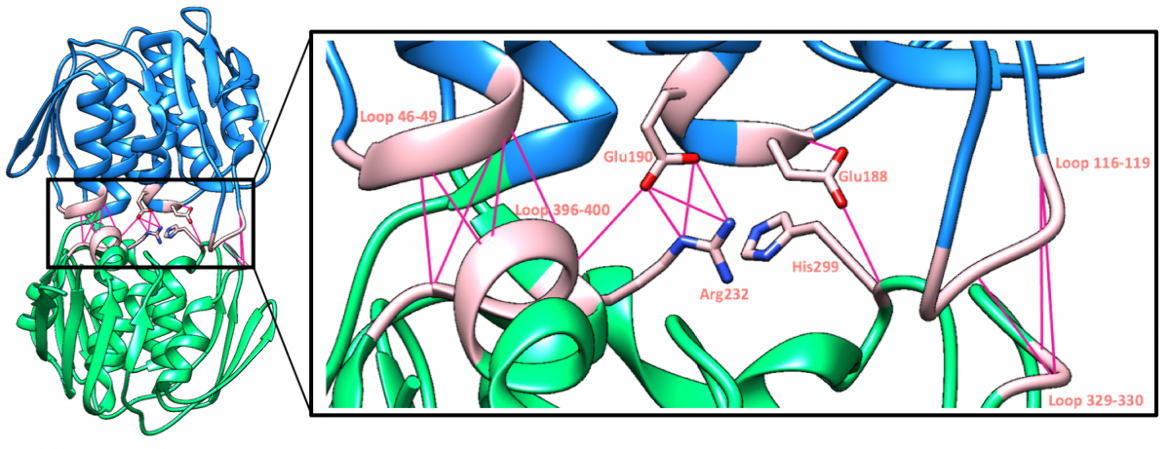
|
| Figure 8. A ribbon representation of the hydrogen bond interactions that occur in the catalytic pocket of MurA (PDB structure: 3upk, Zhu et al., 2012). |
The uridinyl ring of the substrate, UDP-GlcNAc, is sandwiched between the hydrophobic surfaces created by Arg120 and Pro121 on one side and by Leu124 on the other side (Zoeiby et al., 2003), as can be seen in Figure 9. This positioning of the substrate allows for a good fit of the molecular surfaces (Skarzynski et al., 1996). Moreover, some of the main substrate-binding interactions include hydrogen bonds with Arg91, Arg120, Ser162, Val163, and Gly164 (Skarzynski et al., 1996), as shown in Figure 10.
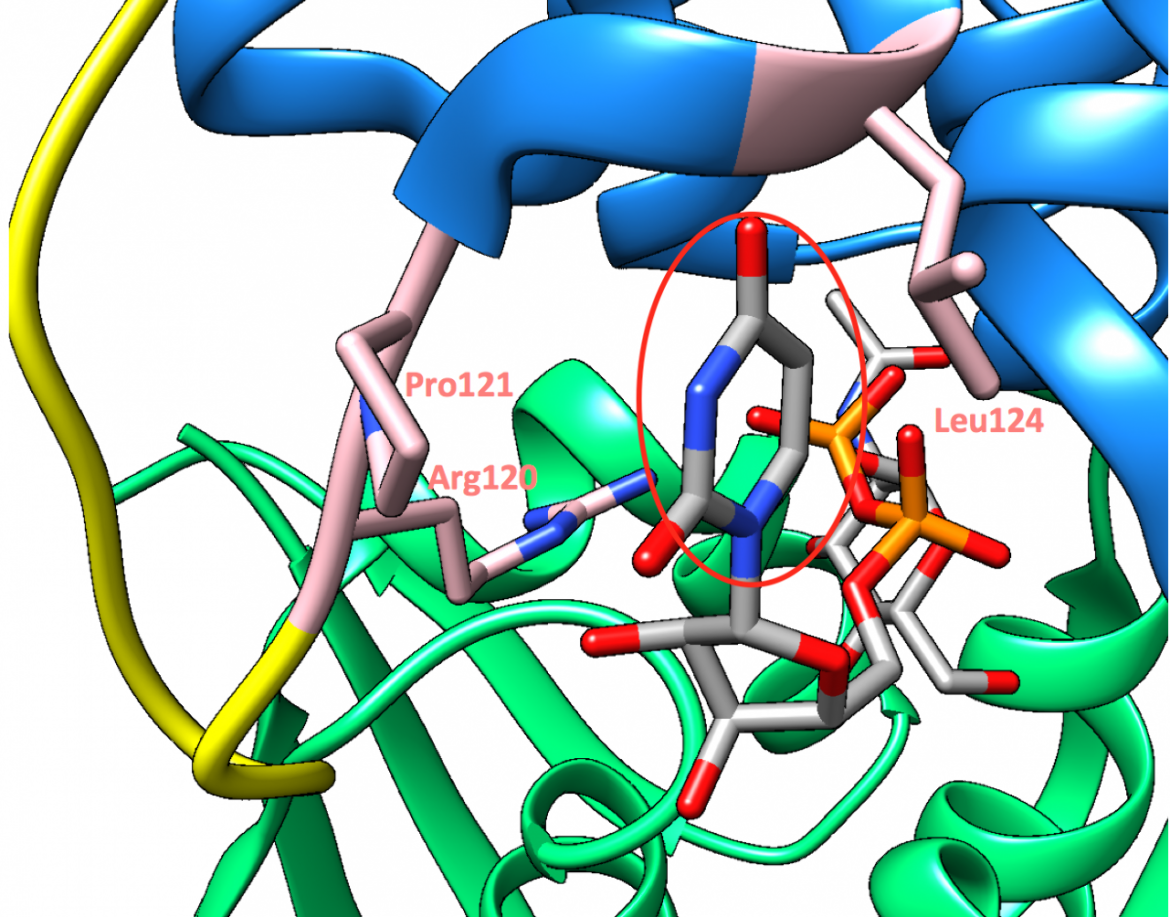
|
| Figure 9. A closeup of the UDP-GlcNAc substrate shows its uridinyl ring (circled in red) sandwiched between Arg120, Pro121, and Leu124 (PDB structure: 3upk, Zhu et al., 2012). |
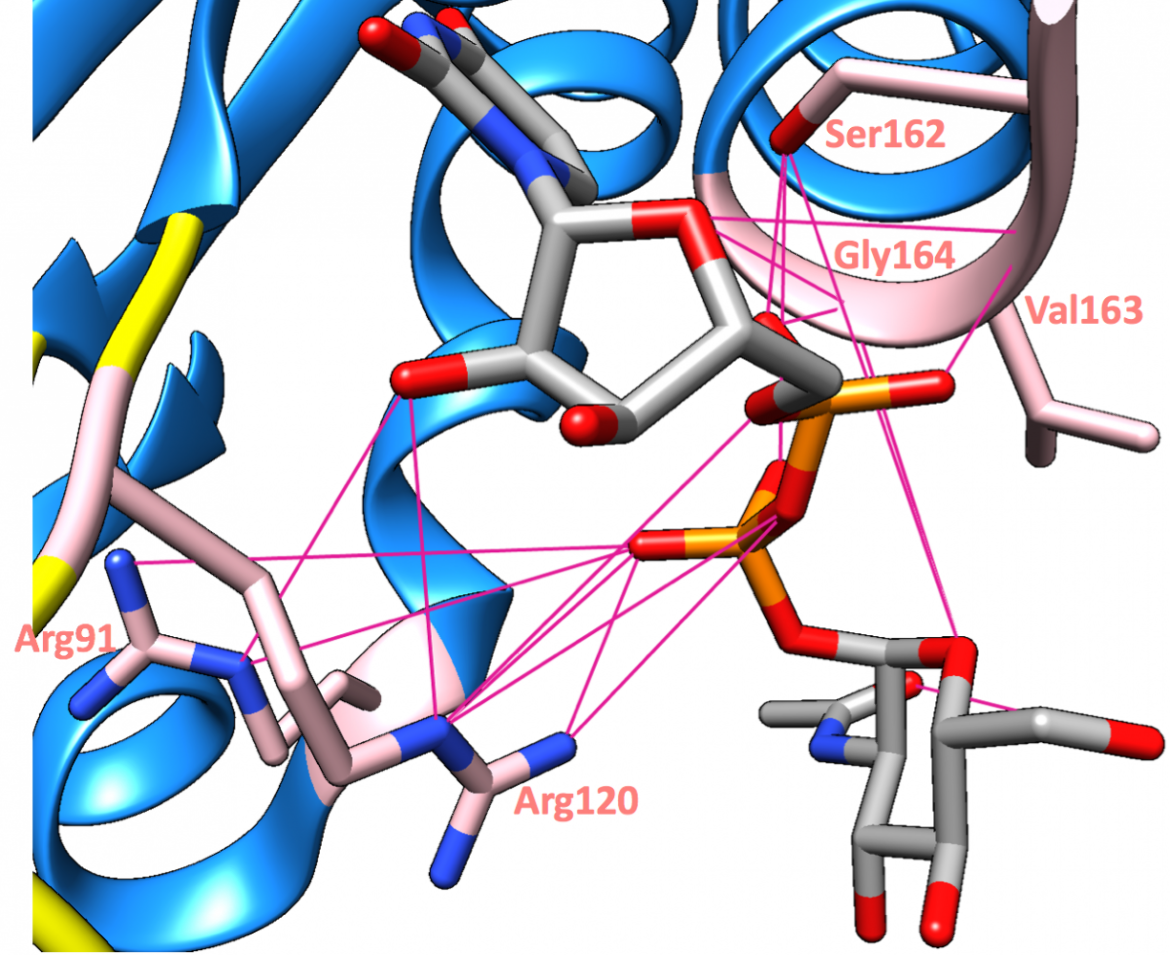
|
| Figure 10. The principal hydrogen bond interactions (magenta lines) made by UDP-GlcNAc are shown (PDB structure: 3upk, Zhu et al., 2012). |
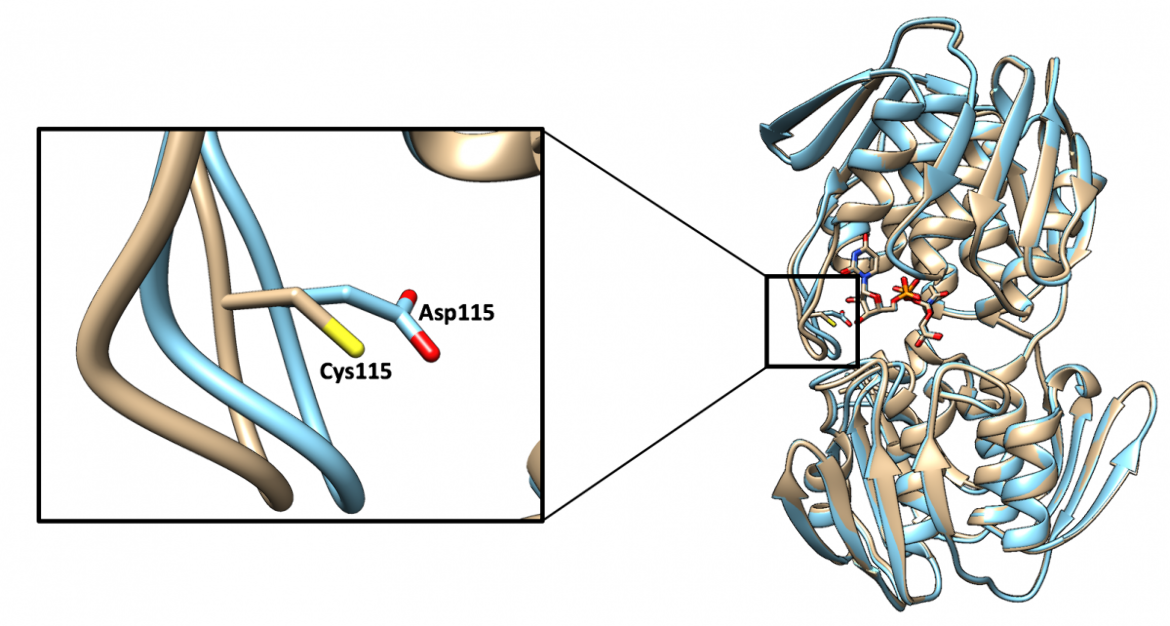
It would normally be expected for an enzyme to become inactive if its catalytic residue were to be mutated; however, in the case of MurA, the enzyme remains highly active when the catalytic Cys115 residue is mutated to an aspartate residue, shown in Figure 11 (Skarzynski et al., 1996). Although this mutation allows the enzyme to retain its function, the new residue can no longer be covalently modified by fosfomycin, making the enzyme resistant to the antibacterial drug (Skarzynski et al., 1996).
As a matter of fact, Mycobacterium tuberculosis MurA has a naturally occurring aspartate residue in the place of a catalytic cysteine residue at position 117, and this allows the bacteria to be intrinsically resistant to fosfomycin; nevertheless, the enzymatic function is not compromised (Jia et al., 2017). These findings indicate that the catalytic cysteine residue performs the role of a general acid in the protonation of PEP during the reaction, a role that can also be fulfilled by an aspartate residue, thus allowing the enzyme to remain active despite the mutation.
Pharmacological Implications
Most bacteria are dependent on a functional MurA enzyme in order to survive because this transferase enzyme catalyzes the first committed step of peptidoglycan biosynthesis (Han et al., 2010). This means that MurA initiates the formation of the nucleotide muramyl peptides that will serve as precursors to the cell wall building blocks (Hendlin et al., 1969). If MurA is inhibited, the cell wall building blocks, and thus the bacterial cell wall, can no longer form, which would lead to bacterial cell lysis and death. Therefore, this enzyme remains an attractive target for novel antibacterial agents.
Off-Target Considerations
The US FDA approved antibacterial drug fosfomycin not only inhibits the bacterial enzyme MurA but also plays a role in the modulation of the host immune system (Falagas et al., 2016). Fosfomycin enhances the ability of neutrophils to phagocytically kill invading pathogens, and it was seen that neutrophils have an enhanced bactericidal ability in the presence of fosfomycin compared to other antimicrobial agents (Falagas et al., 2016).
References
Eschenburg, S., Kabsch, W., Healy, M. L., and Schönbrunn, E. (2003) A New View of the Mechanisms of UDP-N-Acetylglucosamine Enolpyruvyl Transferase (MurA) and 5-Enolpyruvylshikimate-3-phosphate Synthase (AroA) Derived from X-ray Structures of Their Tetrahedral Reaction Intermediate States. Journal of Biological Chemistry 278, 49215-49222. https://doi.org/10.1074/jbc.m309741200
Falagas, M. E., Vouloumanou, E. K., Samonis, G., and Vardakas, K. Z. (2016) Fosfomycin. Clinical Microbiology Reviews 29, 321-347. https://doi.org/10.1128/cmr.00068-15
Han, H., Yang, Y., Olesen, S. H., Becker, A., Betzi, S., and Schönbrunn, E. (2010) The Fungal Product Terreic Acid Is a Covalent Inhibitor of the Bacterial Cell Wall Biosynthetic Enzyme UDP-N-Acetylglucosamine 1-Carboxyvinyltransferase (MurA). Biochemistry 49, 4276-4282. https://doi.org/10.1021/bi100365b PDB structure: 3kr6
Hendlin, D., Stapley, E. O., Jackson, M., Wallick, H., Miller, A. K., Wolf, F. J., Miller, T. W., Chaiet, L., Kahan, F. M., Foltz, E. L., Woodruff, H. B., Mata, J. M., Hernandez, S., and Mochales, S. (1969) Phosphonomycin, a New Antibiotic Produced by Strains of Streptomyces. Science 166, 122-123. https://doi.org/10.1126/science.166.3901.122
Jia, B., Raphenya, A. R., Alcock, B., Waglechner, N., Guo, P., Tsang, K. K., Lago, B. A., Dave, B. M., Pereira, S., Sharma, A. N., Doshi, S., Courtot, M., Lo, R., Williams, L. E., Frye, J. G., Elsayegh, T., Sardar, D. Westman, E. L., Pawlowski, A. C., Johnson, T. A., Brinkman, F. S., Wright, G. D., and McArthur, A. G. (2017) CARD 2017: expansion and model-centric curation of the Comprehensive Antibiotic Resistance Database. Nucleic Acids Research 45, D566-573. https://doi.org/10.1093/nar/gkw1004
Mizyed, S., Oddone, A., Byczynski, B., Hughes, D. W., and Berti, P. J. (2005) UDP-N-acetylmuramic Acid (UDP-MurNAc) Is a Potent Inhibitor of MurA (Enolpyruvyl-UDP-GlcNAc Synthase). Biochemistry 44, 4011-4017. https://doi.org/10.1021/bi047704w
Samland, A. K., Amrhein, N., and Macheroux, P. (1999) Lysine 22 in UDP-N-Acetylglucosamine Enolpyruvyl Transferase from Enterobacter cloacae Is Crucial for Enzymatic Activity and the Formation of Covalent Adducts with the Substrate Phosphoenolpyruvate and the Antibiotic Fosfomycin. Biochemistry 38, 13162-13169. https://doi.org/10.1021/bi991041e
Schönbrunn, E., Sack, S., Eschenburg, S., Perrakis, A., Krekel, F., Amrhein, N., and Mandelkow, E. (1996) Crystal structure of UDP-N-acetylglucosamine enolpyruvyltransferase, the target of the antibiotic fosfomycin. Structure 4, 1065-1075. https://doi.org/10.1016/s0969-2126(96)00113-x
Skarzynski, T., Mistry, A., Wonacott, A., Hutchinson, S. E., Kelly, V. A., and Duncan, K. (1996) Structure of UDP-N-acetylglucosamine enolpyruvyl transferase, an enzyme essential for the synthesis of bacterial peptidoglycan, complexed with substrate UDP-N-acetylglucosamine and the drug fosfomycin. Structure 4, 1465-1474. https://doi.org/10.1016/s0969-2126(96)00153-0 PDB structure: 1uae
Zhu, J., Yang, Y., Han, H., Betzi, S., Olesen, S. H., Marsilio, F., and Schönbrunn, E. (2012) Functional Consequence of Covalent Reaction of Phosphoenolpyruvate with UDP-N-acetylglucosamine 1-Carboxyvinyltransferase (MurA). Journal of Biological Chemistry 287, 12657-12667. https://doi.org/10.1074/jbc.m112.342725 PDB structures: 3spb, 3upk, 3swa, 3su9, 3v4t
Zoeiby, A. E., Sanschagrin, F., and Levesque, R. C. (2002) Structure and function of the Mur enzymes: development of novel inhibitors. Molecular Microbiology 47, 1-12. https://doi.org/10.1046/j.1365-2958.2003.03289.x
March 2025, Gauri Patel: Reviewed by Dr. Lynn Silver
https://doi.org/10.2210/rcsb_pdb/GH/AMR/drugs/antibiotics/cellwall-biosynth/mura