Molecule of the Month: Glucansucrase
Bacteria adhere to our teeth by building sticky sugar chains
Sticky Sugars
Growing Glucans

The Same, Only Different
Exploring the Structure
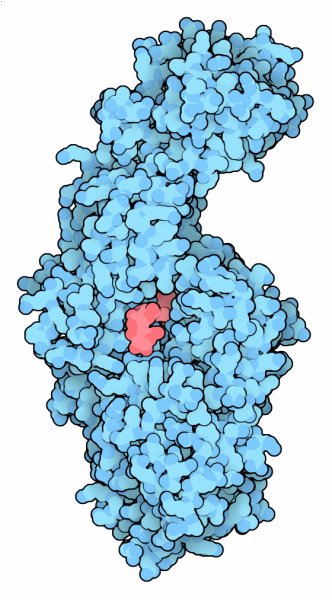
Glucansucrase (PDB entries 3hz3 and 3kll)

Structures have been solved for glucansucrase from two different bacteria: Streptococcus mutans, shown in the previous sections, and Lactobacillus reuteri, shown in the Jmol. The Lactobacillus enzyme has been studied in several ways. First, a mutant form was used to reveal the structure of the complex with sucrose; the mutation hobbles the enzyme so that it can't perform the cleavage reaction (PDB entry 3hz3 ). Also, a complex with maltose was solved to get a view of how larger sugar chains may bind (PDB entry 3kll ). The image here combines both structures to simulate how the enzyme might look in the middle of the reaction, when fructose is cleaved away, leaving a glucose ready to add to the glucan chain. Click on the image to see the two structures and this simulated complex.
Topics for Further Discussion
- When you look at the complexes of glucansucrases with sugars, you will find that there are several sites outside of the active site where additional sugar molecules bind. Do you think that these may play a role in the function of the protein, or are they just artifacts of the experiment?
- You can use the "Compare Structures" tool to explore the circular permutation of glucansucrases and alpha-amylase.
Related PDB-101 Resources
- Browse You and Your Health
- Browse Enzymes
References
- K. Ito, S. Ito, T. Shimamura, S. Weyand, Y. Kawarasaki, T. Misaka, K. Abe, T. Kobayashi, A. D. Cameron and S. Iwata (2011) Crystal structure of glucansucrase from the dental caries pathogen Streptococcus mutans. Journal of Molecular Biology 408, 177-186.
- A. Vujicic-Zagar, T. Pijning, S. Kralj, C. A. Lopez, W. Eeuwema, L. Dijkhuizen and B. W. Dijkstra (2010) Crystal structure of a 117 kDa glucansucrase fragment provides insight into evolution and product specificity of GH70 enzymes. Proceedings of the National Academy of Sciences USA 107, 21406-21411.
- J. D. B. Featherstone (2008) Dental caries: a dynamic disease process. Australian Dental Journal 53, 286-291.
June 2011, David Goodsell
http://doi.org/10.2210/rcsb_pdb/mom_2011_6


